| Oberlin College Biology Department | |
 |
|
Mike MooreProfessor of BiologyOberlin CollegeScience Center K111 (440) 775-6876 Ph.D. University of Texas, 2005 M.S. University of Illinois, 1999 B.S. College of William and Mary, 1995 Publications (also visit my Google Scholar page) |
 Follow me on Twitter: @gypsumbotany Follow me on Twitter: @gypsumbotany |
Moore Lab Research: Plant Systematics
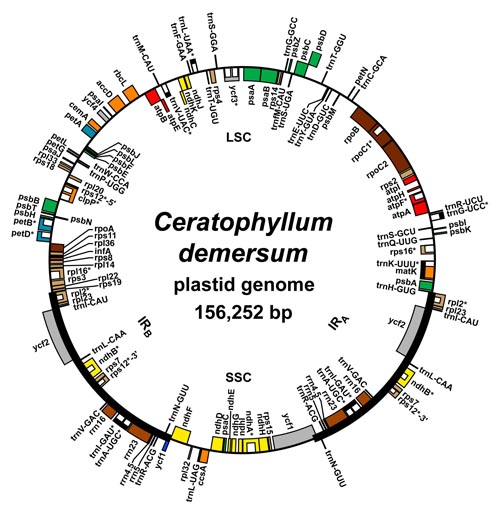
Research in my lab lies within the field of plant systematics—the study of the origin and evolution of plant diversity. We are interested in exploring various problems in flowering plant evolution using molecular phylogenetic and phylogeographic approaches. In other words, we reconstruct the evolutionary relationships (phylogeny) among plants by generating and analyzing DNA sequence data for various gene regions. We use these molecular phylogenies to test hypotheses about morphological, ecological, and molecular evolution within the plant groups we are interested in. For example, one of our major current interests is understanding how unusual soils have interacted with past climate changes to influence the diversification of plants in the desert Southwest. This project involves extensive fieldwork to collect leaf samples, which we bring back to Oberlin for molecular analyses. For more on this project, see below.
Systematics incorporates ideas and techniques from across biology, including fields like evolutionary biology, anatomy, ecology, molecular biology, biogeography, and bioinformatics. Consequently, students in my lab have the opportunity to do fieldwork, labwork, and computational work.
Moore Lab research featured in the video series "Plants Are Cool, Too!"—watch below or read the associated news article:
Watch the following clip to learn a bit more about my research and teaching philosphy:
Research in my lab is inherently computational, whether it is reconstructing evolutionary trees using phylogenetic algorithms or assembling chloroplast genomes using millions of short DNA sequences generated from next-generation sequencers. The Oberlin supercomputer plays a key role in these analyses. Recently I was part of a team of Oberlin faculty (including Manish Mehta and Matt Elrod of the Chemistry Dept., Rob Owen of the Physics Dept., and Aaron Goldman of the Biology Dept.) that obtained a $500,000 grant from the National Science Foundation to purchase and install a vastly improved high-performance computing cluster. For more on this exciting opportunity, read this brief feature from the Oberlin News Center.
And finally—and most importantly—my research would not be possible without the many talented Oberlin students I have mentored over the years. Working with Oberlin undergraduates is the most enjoyable part of my job. Below are some photos of folks who have worked in my lab over the years, including many Obies.
 |
The Moore lab gang, Spring semester 2015. (In the tree, left to right) Erin Johnson, Vera Hutchison, Noah Last; (Front row, left to right) Jill Sarazen, Zoë Feder, Ellen Drake. |
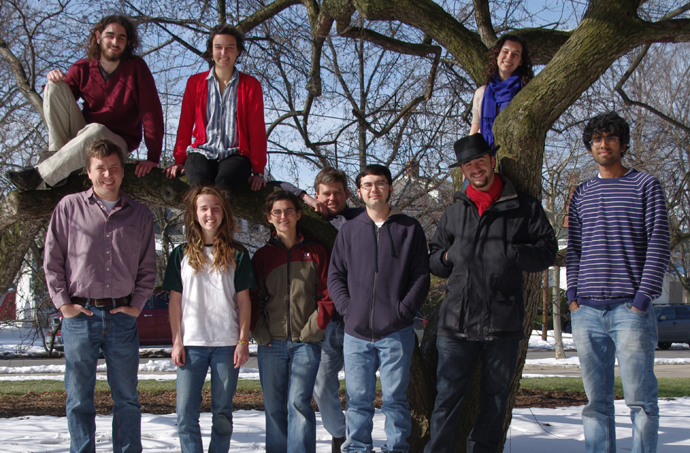 |
The Moore lab crew, Spring semester 2013. (In the tree, left to right) Jonah Joffe, Lila Leatherman, Rebecca Mostow; (Front row, left to right) Mike Moore, Emma Ray, Arianna Goodman, Norm Douglas, James Medina, Spencer Wight, Shiva Mandala. |
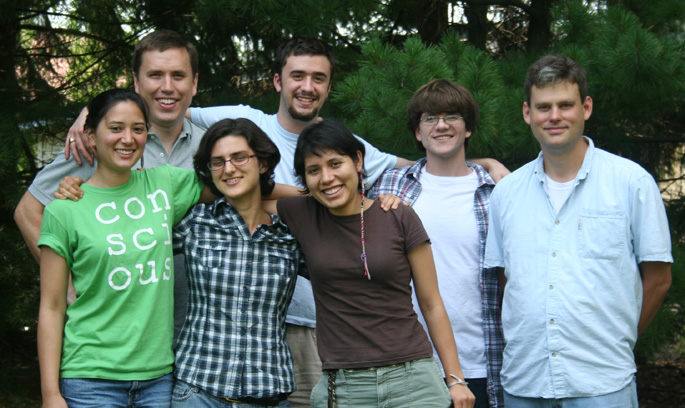 |
| The Moore lab summer 2012 research gang. (Back row, left to right) Mike Moore, Spencer Wight, Trip Freeburg, Norm Douglas; (Front row, left to right) Chloe Drummond, Arianna Goodman, Nidia Mendoza Díaz. |
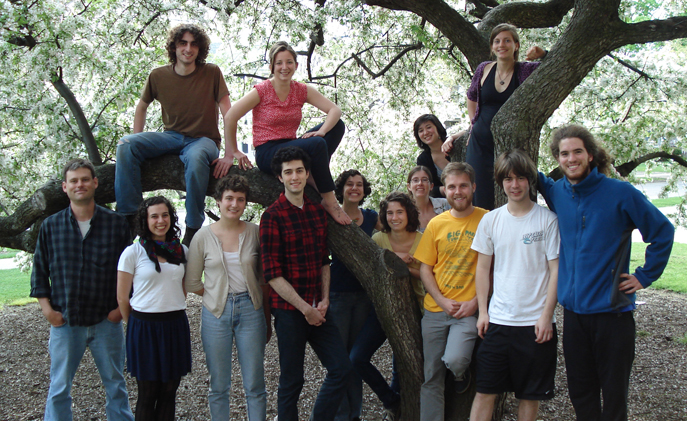 |
Moore Lab, Spring 2011. L to R: (Back row) Jonah Joffe, Heather-Rose Kates, Chloe Drummond, Sophia Weinmann; (Front row) Norm Douglas, Rebecca Mostow, Julia Ruby, Gabe Stalberg, Riva Bruenn, Rachel Plumb, Carolyn Stange, Matt Croley, Liam Sharninghausen, Chris Canning. Not pictured: Kate Hampilos, Flora Samis. |
My lab is full of opportunities for the motivated student. Students in the Moore Lab learn many techniques, ranging from DNA isolation to sequence editing/analysis to cloning. In addition to lab work, in summers we often go on expeditions to the desert Southwest to collect plants for our research.
Right: Anna Brunner and Jeffrey Sanders collect gypsum endemic plants at Sevilleta National Wildlife Refuge, New Mexico. |
 |
 |
 |
Joey Charboneau, Anna Brunner, Jeffrey Sanders, and Mike Moore at White Sands National Monument, New Mexico |
Mike Moore, Anna Brunner, Heather-Rose Kates, and Jeffrey Sanders in Valley of the Gods, southeastern Utah |
Research in my lab currently focuses on three broad areas:
(1) The Origin and Evolution of Gypsum Endemic Plants in the Chihuahuan Desert
To learn more about my research on gypsum endemism, click here or on the image below:
(2) The evolution of breeding systems in Hawaiian plants
My collaborators and I are reconstructing the evolutionary history as well as patterns of gene flow in the Hawaiian endemic genus Schiedea, one of the most diverse plant groups in the Hawaiian islands.
(3) Flowering Plant Phylogenomics
To learn more about my efforts to resolve angiosperm phylogeny, click here or on the image:
Publications (also see my Google Scholar page):
(*Oberlin undergraduate authors from Moore lab)
Zhang, Y., T. Deng, L. Sun, J. B. Landis, M. J. Moore, H. Wang, Y. Wang, X. Hao, J. Chen, S. Li, M. Xu, P.-T. Puno, P. H. Raven, and H. Sun. In press. Phylogenetic patterns and signal in secondary metabolites across the seed plant tree of life. National Science Review.
Zhang, R., Y.-H. Wang, J.-J. Jin, A. Bruneau, D. Cardoso, L. Paganucci de Queiroz, M. J. Moore, S.-D. Zhang, S.-Y. Chen, J. Wang, D.-Z. Li, and T.-S. Yi. In press. Plastid phylogenomic insights into deep relationships of Leguminosae with conflicting signals on some nodes. Systematic Biology.
da Cunho Neto, I. L., M. R. Pace, N. A. Douglas, M. H. Nee, C. Farney C. de Sá, M. J. Moore, and V. Angyalossy. In press. Medullary bundles in Nyctaginaceae: their diversity, distribution, development and evolution. American Journal of Botany.
Wang, H.-X., M. J. Moore, R. L. Barrett, S. Landrein, S. Sakaguchi, M. Maki, J. Wen, and H. Wang. In press. Plastome phylogenomic insight into the Sino-Japanese biogeography of Diabelia (Caprifoliaceae). Journal of Systematics and Evolution Early View. [HTML]
Ochoterena, H., H. Flores, C. Gómez, and M. J. Moore. In press. Gypsum and plant species: a wonder of Cuatro Ciénegas, Coahuila. Part of the book series: Cuatro Ciénegas Basin: An Endangered Hyperdiverse Oasis. Springer.
Mendoza-Díaz, N., H. Ochoterena, Michael J. Moore, and H. Flores-Olvera. 2020. Molecular and morphological evidence reveals a new species of Antiphytum (Echiochiloideae, Boraginaceae) from Guerrero, Mexico. Systematic Botany 45(1): 190-199. [HTML]
Zhu, Z., J.-H. Wang, S. Sakaguchi, K. K. Zhao, M. J. Moore, and H. Wang. 2020. The complete plastome sequences of two Neottia species and comparative analysis with other Neottieae species (Orchidaceae). Folia Geobotanica. [HTML]
Wang, J-H., M. J. Moore, K.-K. Zhao, S. Landrein, H.-X. Wang, Z.-X. Zhu, S. K. Sahu, B. Liu, H. Liu, and H.-F. Wang. 2020. Plastid phylogenomic insights into the evolution of the Caprifoliaceae s.l. (Dipsacales). Molecular Phylogenetics and Evolution 142: 106641. [HTML]
Sheehan, H., T. Feng, N. Walker-Hale, S. Lopez-Nieves, B. Pucker, R. Guo, W. C. Yim, R. Badagami, A. Timoneda, L. Zhao, *H. Tiley, D. Copetti, M. Sanderson, J. Cushman, M. J. Moore, S. A. Smith, and S. F. Brockington. 2019. Evolution of L-DOPA 4,5-dioxygenase activity allows for recurrent specialisation to betalain pigmentation in Caryophyllales. New Phytologist Early View. [HTML]
Zhang, R., J. Jin, M. J. Moore, and T. Yang. 2019. Assembly and comparative analyses of the mitochondrial genome of Castanospermum australe A.Cunn. & C.Fraser (Papilionoideae, Leguminosae). Australian Systematic Botany 32(6): 484-494. [HTML]
Zhang, X., T. Deng, M. J. Moore, H. Wang, Z. Li, N. Lin, H. Zhang, A. Meng, H. Wang, Y. Sun, and H. Sun. 2019. Plastome phylogenomics of Saussurea (Asteraceae: Cardueae). BMC Plant Biology 19: 290. [HTML]
Huang, X., T. Deng, M. J. Moore, H. Wang, Z. Li, N. Lin, Z. Yusupov, K. S. Tojibaev, Y. Wang, and H. Sun. 2019. Tropical Asian origin, boreotropical migration and long-distance dispersal in nettles (Urticeae, Urticaceae). Molecular Phylogenetics and Evolution 137: 190-199. [HTML]
Qu, X.-J., M. J. Moore, D.-Z. Li, and T.-S. Yi. 2019. PGA: a software package for rapid, accurate, and flexible batch annotation of plaastomes. Plant Methods 15: 50. [HTML]
Gang, Y., J. Jian, H.-T. Li, J.-B. Yang, *V. S. Mandala, *M. Croley, *R. Mostow, N. Douglas, M. W. Chase, M. Christenhusz, D. E. Soltis, P. S. Soltis, S. A. Smith, S. F. Brockington, M. J. Moore, T.-S. Yi, and D.-Z. Li. 2019. Plastid phylogenomic insights into the evolution of Caryophyllales. Molecular Phylogenetics and Evolution 134: 74-86. [HTML]
Wang, N., Y. Yang, M. J. Moore, S. F. Brockington, J. F. Walker, J. W. Brown, B. Liang, T. Feng, *C. Edwards, *J. Mikenas, *J. Olivieri, *V. Hutchison, A. Timoneda, T. Stoughton, R. Puente, L. Majure, U. Eggli, and S. A. Smith. 2019. Evolution of Portulacineae marked by gene tree conflict and gene family expansion associated with adaptation to harsh environments. Molecular Biology and Evolution 36(1): 112-126. [HTML]
Mendoza-Díaz, N., H. Flores-Olvera, M. G. Simpson, and M. J. Moore. 2018. A new and unusual endemic species from the Chihuahuan Desert, Mexico: Antiphytum geoffreyi (Boraginaceae, Echiochiloideae). Phytotaxa 367(3): 275-283. [HTML]
Sun, Y., M. J. Moore, J. B. Landis, N. Lin, L. Chen, T. Deng, J. Zhang, A. Meng, S. Zhang, K. S. Tojibaev, H. Sun, and H. Wang. 2018. Plastome phylogenomics of the early-diverging eudicot family Berberidaceae. Molecular Phylogenetics and Evolution 128: 203-211. [HTML]
Zhang, H., Jin, J., M. J. Moore, T. Yi, and D. Li. 2018. Plastome characteristics of Cannabaceae. Plant Diversity 40(3): 127-137. [HTML]
Lin, N., T. Deng, M. J. Moore, Y. Sun, W. Sun, D. Luo, H. Wang, J. Zhang, and H. Sun. 2018. Phylogeography of Parasyncalathium souliei (Asteraceae) and its potential application in delimiting phylogeoregions in the Qinghai-Tibet Plateau (QTP) - Hengduan Mountains (HDM) hotspot. Frontiers in Genetics 9: 171. [HTML]
Walker, J. F., Y. Yang, T. Feng, A. Timoneda, *J. Mikenas, *V. Hutchinson, *C. Edwards, N. Wang, S. Ahluwalia, *J. Olivieri, N. Walker-Hale, L. C. Majure, R. Puente, G. Kadereit, M. Lauterbach, U. Eggli, H. Flores Olvera, H. Ochoterena, S. F. Brockington, M. J. Moore, and S. A. Smith. 2018. From cacti to carnivores: Improved phylotranscriptomic sampling and hierarchical homology inference provide further insight to the evolution of Caryophyllales. American Journal of Botany (special issue: Using and Navigating the Plant Tree of Life) 105(3): 446-462. [HTML]
Soltis, D. E., M. J. Moore, E. B. Sessa, S. A. Smith, and P. S. Soltis. 2018. Using and navigating the plant tree of life. American Journal of Botany (special issue: Using and Navigating the Plant Tree of Life) 105(3): 287-290. [HTML]
Lin, N., M. J. Moore, T. Deng, H. Sun, L. Yang, Y. Sun, and H. Wang. 2018. Complete plastome sequencing from Toona (Meliaceae) and phylogenomic analyses within Sapindales. Applications in Plant Sciences 6(4): e01040. [HTML]
Zhu, Z., J. Wang, Y. Cai, K. Zhao, M. J. Moore, and H. Wang. 2018. Complete plastome sequence of Erythropalum scandens (Erythropalaceae), an edible and medicinally important liana in China. Mitochondrial DNA Part B: Resources 3(1): 139-140. [HTML]
Yan, M., P. W. Fritsch, M. J. Moore, T. Feng, A. Meng, J. Yang, T. Deng, C. Zhao, X. Yao, H. Sun, and H. Wang. 2018. Plastome phylogenomics resolves infrafamilial relationships of the Styracaceae and sheds light on the backbone relationships of the Ericales. Molecular Phylogenetics and Evolution 121: 198-211. [HTML]
Smith, S. A., J. W. Brown, Y. Yang, *R. Bruenn, *C. P. Drummond, S. F. Brockington, J. F. Walker, *N. Last, N. A. Douglas, and M. J. Moore. 2018. Disparity, diversity, and duplications in the Caryophyllales. New Phytologist 217(2): 836-854. [HTML]
Yang, Y., M. J. Moore, S. F. Brockington, *J. Mikenas, *J. Olivieri, J. F. Walker, and S. A. Smith. 2018. Improved transcriptome sampling pinpoints 26 ancient and more recent paleopolyploidy events in Caryophyllales, including two allopolyploidy events. New Phytologist 217(2): 855-870. [HTML]
Muller, C. T., M. J. Moore, *Z. Feder, *H. Tiley, and R. Drenovsky. 2017. Phylogenetic patterns of foliar mineral nutrient accumulation among gypsophilic plants and their relatives in the Chihuahuan Desert. American Journal of Botany 104(10): 1442-1450. [HTML]
Guo, R., J. B. Landis, M. J. Moore, A. Meng, S. Jian, X. Yao, and H. Wang. 2017. Development and application of transcriptome-derived microsatellites in Actinidia eriantha (Actinidiaceae). Frontiers in Plant Science 8: 1383. [HTML]
Sun, Y., M. J. Moore, N. Lin, J. Li, and H. Wang. 2017. Complete plastome sequencing of both living species of Circaeasteraceae reveals unusual rearrangements and the loss of the ndh gene family. BMC Genomics 18: 592. [HTML]
Walker, J. F., Y. Yang, M. J. Moore, *J. Mikenas, A. Timoneda, S. F. Brockington, and S. A. Smith. 2017. Widespread paleopolyploidy, gene tree conflict, and recalcitrant relationships among the carnivorous Caryophyllales. American Journal of Botany 104(6): 858-867. [HTML]
Yang, Y., M. J. Moore, S. F. Brockington, A. Timoneda-Monfort, T. Feng, H. E. Marx, J. F. Walker, and S. A. Smith. 2017. An efficient field and laboratory workflow for plant phylotranscriptomic projects. Applications in Plant Sciences 5(3): 1600128. [HTML]
Feng, T., M. J. Moore, M. Yan, Y. Sun, A. Meng, X. Li, J. Li, and H. Wang. In press. Phylogenetic study of the tribe Potentilleae (Rosaceae), with further insight into the disintegration of Sibbaldia. Journal of Systematics and Evolution. 55(3): 177-191. [HTML]
Yan, M., M. J. Moore, A. Meng, X. Yao, and H. Wang. 2017. The first complete plastome sequence of the basal asterid family Styracaceae (Ericales) reveals a large inversion. Plant Systematics and Evolution 303(1): 61-70. [HTML]
Sun, Y., M. J. Moore, S. Zhang, P. S. Soltis, D. E. Soltis, A. Meng, J. Li, and H. Wang. 2016. Phylogenomic and structural analyses of 18 complete plastomes across nearly all families of early-diverging eudicots, including an angiosperm-wide analysis of IR gene content evolution. Molecular Phylogenetics and Evolution 96(3): 93-101. [HTML] [PDF]
Lewis, E. M., J. B. Fant, M. J. Moore, A. P. Hastings, E. L. Larson, A. Agrawal, and K. A. Skogen. 2016. Microsatellites for Oenothera gayleana and O. hartwegii subsp. filifolia (Onagraceae), and their utility in section Calylophus. Applications in Plant Sciences 4(2): 1500107. [HTML] [PDF]
Kim, H. T., J. S. Kim, M. J. Moore, K. M. Neubig, N. H. Williams, W. M. Whitten, and J. H. Kim. 2015. Seven new complete plastome sequences reveal rampant independent loss of the ndh gene family across orchids and associated instability of the Inverted Repeat/Small Single-Copy region boundaries. PLoS ONE 10(11): e0142215. [HTML] [PDF]
Smith, S. A., M. J. Moore, J. Brown, and Y. Yang. 2015. Analysis of phylogenomic datasets reveals conflict, concordance, and gene duplications, with examples from animals and plants. BMC Evolutionary Biology 15: 150. [HTML] [PDF]
Brockington, S. F., Y. Yang, F. Gandia-Herrero, S. Covshoff, J. M. Hibberd, R. F. Sage, G. K.-S. Wong, M. J. Moore, and S. A. Smith. 2015. Lineage-specific gene radiations underlie the evolution of novel betalain pigmentation in Caryophyllales. New Phytologist 207(4): 1170-1180. [HTML] [PDF]
Yang, Y., M. J. Moore, S. F. Brockington, D. E. Soltis, G. K.-S. Wong, E. J. Carpenter, Y. Zhang, L. Chen, Z. Yan, Y. Xie, R. F. Sage, S. Covshoff, J. M. Hibberd, M. N. Nelson, and S. A. Smith. 2015. Dissecting molecular evolution in the highly diverse plant clade Caryophyllales using transcriptome sequencing. Molecular Biology and Evolution 32(8): 2001-2014. [HTML] [PDF]
Feng, T., M. J. Moore, Y. Sun, A. Meng, H. Chu, J. Li, and H. Wang. 2015. A new species of Argentina (Rosaceae, Potentilleae) from Southeast Tibet, with reference to the taxonomic status of the genus. Plant Systematics and Evolution 301(3): 911-921 [HTML] [PDF]
Alexander, P. J., N. A. Douglas, H. Ochoterena, H. Flores Olvera, and M. J. Moore. 2014. Recent findings on the gypsum flora of The Rim of the Guadalupe Mountains, New Mexico: a new species of Nerisyrenia, a new state record, and an updated checklist. Journal of the Botanical Research Institute of Texas 8(2): 383-394. [PDF]
Moore, M. J., J. F. Mota, N. A. Douglas, H. Flores Olvera, and H. Ochoterena. 2014. The ecology, assembly, and evolution of gypsophile floras. Pp. 97- 128 in Plant Ecology and Evolution in Harsh Environments, Eds. N. Rajakaruna, R. Boyd, and T. Harris. Nova Science Publishers: Hauppauge, NY. [PDF]
Straub, S. C. K., M. J. Moore, P. S. Soltis, D. E. Soltis, A. Liston, and T. Livshultz. 2014. Phylogenetic signal detection from an ancient rapid radiation: Effects of noise reduction, long-branch attraction, and model selection in crown clade Apocynaceae. Molecular Phylogenetics and Evolution 80: 169-185. [HTML] [PDF]
Turner, B. L. and M. J. Moore. 2014. Oenothera gayleana (Oenothera sect. Calylophus, Onagraceae), a new gypsophile from Texas, New Mexico, and Oklahoma. Phytologia 96(3): 200-206. [PDF]
Sun, Y., M J. Moore, L. Yue, T. Feng, H. Chu, S. Chen, Y. Ji, H. Wang, and J. Li. 2014. Chloroplast phylogeography of the East Asian Arcto-Tertiary relict Tetracentron sinense (Trochodendraceae). Journal of Biogeography 41(9): 1721-1732. [HTML] [PDF]
Drew, B. T., B. R. Ruhfel, S. A. Smith, M. J. Moore, B. G. Briggs, M. A. Gitzendanner, P. S. Soltis, and D. E. Soltis. 2014. Another look at the root of the angiosperms reveals a familiar tale. Systematic Biology 63(3): 368-382. [HTML] [PDF]
Uribe-Convers, S., J. R. Duke, M. J. Moore, and D. C. Tank. 2014. A long PCR-based approach for DNA enrichment prior to next-generation sequencing for systematic studies. Applications in Plant Sciences 2(1): 1300063. [HTML] [PDF]
Sun, Y. X., M. J. Moore, P. S. Soltis, D. E. Soltis, J. Q. Li, and H. C. Wang. 2013. Complete plastid genome sequencing of Trochodendraceae reveals a significant expansion of the Inverted Repeat and suggests a Paleogene divergence between the two extant species. PLoS ONE 8(4): e60429. [HTML] [PDF]
Stull, G. W., M. J. Moore, *V. S. Mandala, N. A. Douglas, H.-R. Kates, X. Qi, S. F. Brockington, P. S. Soltis, D. E. Soltis, and M. A. Gitzendanner. 2013. A targeted enrichment strategy for massively parallel sequencing of angiosperm plastid genomes. Applications in Plant Sciences 1(2): 1-7. [HTML] [PDF]
Arakaki, M., P.-A. Christin, R. Nyffeler, A. Lendel, U. Eggli, R. M. Ogburn, E. Spriggs, M. J. Moore, and E. J. Edwards. 2011. Contemporaneous and recent radiations of the world's major succulent plant lineages. Proceedings of the National Academy of Sciences USA 108(20): 8379-8384. [HTML] [PDF]
Moore, M. J., N. Hassan, M. A. Gitzendanner, *R. A. Bruenn, *M. Croley, A. Vandeventer, J. W. Horn, A. Dhingra, S. F. Brockington, M. Latvis, J. Ramdial, R. Alexandre, A. Piedrahita, Z. Xi, C. C. Davis, P. S. Soltis, and D. E. Soltis. 2011. Phylogenetic analysis of the plastid inverted repeat for 244 species: insights into deeper-level angiosperm relationships from a long, slowly evolving sequence region. International Journal of Plant Sciences 172(4): 541-558. [HTML] [PDF]
Soltis, D. E., S. A. Smith, N. Cellinese, K. J. Wurdack, D. C. Tank, S. F. Brockington, N. F. Refulio-Rodriguez, J. B. Walker, M. J. Moore, B. S. Carlsward, C. D. Bell, M. Latvis, S. Crawley, C. Black, D. Diouf, Z. Xi, M. A. Gitzendanner, K. J. Sytsma, Y.-L. Qiu, K. W. Hilu, C. C. Davis, M. J. Sanderson, R. G. Olmstead, W. S. Judd, M. J. Donoghue, and P. S. Soltis. 2011. Angiosperm phylogeny: 17 genes, 640 taxa. American Journal of Botany 98(4): 704-730. [HTML] [PDF]
Moore, M. J., P. S. Soltis, C. D. Bell, J. G. Burleight, and D. E. Soltis. 2010. Phylogenetic analysis of 83 plastid genes further resolves the early diversification of eudicots. Proceedings of the National Academy of Sciences USA 107(10): 4623-4628. [HTML] [PDF] (Open Access)
Soltis, D. E., M. J. Moore, J. G. Burleigh, C. D. Bell, and P. S. Soltis. 2010. Assembling the angiosperm tree of life: progress and future prospects. Annals of the Missouri Botanical Garden 97(4): 514-526. [HTML] [PDF]
Givnish, T. J., M. Ames, J. R. McNeal, M. R. McKain, P. R. Steele, C. W. dePamphilis, S. W. Graham, J. C. Pires, D. W. Stevenson, W. B. Zomlefer, B. G. Briggs, M. R. Duvall, M. J. Moore, J. M. Heaney, D. E. Soltis, P. S. Soltis, K. Thiele, and J. H. Leebens-Mack. 2010. Assembling the tree of the monocotyledons: Plastome sequence phylogeny and evolution of Poales. Annals of the Missouri Botanical Garden 97(4): 584-616. [HTML] [PDF]
Soltis, D. E., G. Burleigh, W.B. Barbazuk, M. J. Moore, and P. S. Soltis. 2010. Advances in the use of next-generation sequence data in plant systematics and evolution. ISHS Acta Horticulturae 859 (International Symposium on Molecular Markers in Horticulture). [link]
Brockington, S. F., R. Alexandre, J. Ramdial, M. J. Moore, S. Crawley, A. Dhingra, K. Hilu, P. S. Soltis, and D. E. Soltis. 2009. Phylogeny of the Caryophyllales sensu lato: Revisiting hypotheses on pollination biology and perianth differentiation in the core Caryophyllales. International Journal of Plant Sciences 170(5): 627-643. [HTML] [PDF]
Soltis, D. E., M. J. Moore, J. G. Burleigh, and P. S. Soltis 2009. Molecular markers and concepts of plant evolutionary relationships: progress, promise, and future prospects. Critical Reviews in Plant Sciences 28(1-2): 1-15. [HTML/PDF]
Wang, H., M. J. Moore, P. S. Soltis, C. D. Bell, S. F. Brockington, R. Alexandre, C. C. Davis, M. Latvis, S. R. Manchester, and D. E. Soltis. 2009. Rosid radiation and the rapid rise of angiosperm-dominated forests. Proceedings of the National Academy of Sciences USA 106(10): 3853-3858. [HTML] [PDF]
Soltis, P. S., S. F. Brockington, M.-J. Yoo, A. Piedrahita, M. Latvis, M. J. Moore, A. S. Chanderbali, and D. E. Soltis. 2009. Floral variation and floral genetics in basal angiosperms. American Journal of Botany 96(1): 110-128. [HTML] [PDF]
Neubig, K. M., W. M. Whitten, B. S. Carlsward, M. A. Blanco, L. Endara, N. H. Williams, and M. J. Moore. 2009. Phylogenetic utility of ycf1 in orchids: a plastid gene more variable than matK. Plant Systematics and Evolution 277 (1-2): 75-84. [HTML] [PDF]
Moore, M. J. 2009. Angiosperm phylogenetics. McGraw-Hill 2009 Yearbook of Science & Technology. McGraw-Hill, New York. [HTML]
Jian, S., P. S. Soltis, M. A. Gitzendanner, M. J. Moore, R. Li, T. A. Hendry, Y.-L. Qiu, A. Dhingra, C. D. Bell, and D. E. Soltis. 2008. Resolving an ancient, rapid radiation in Saxifragales. Systematic Biology 57(1): 38-57. [HTML] [PDF]
Moore, M. J., C. D. Bell, P. S. Soltis, and D. E. Soltis. 2007. Using plastid genome-scale data to resolve enigmatic relationships among basal angiosperms. Proceedings of the National Academy of Sciences USA 104(49): 19363-19368. [HTML] [PDF]
Moore, M. J., and R. K. Jansen. 2007. Origins and biogeography of gypsophily in the Chihuahuan Desert plant group Tiquilia subg. Eddya (Boraginaceae). Systematic Botany 32(2): 392-414. [HTML] [PDF]
Moore, M. J., A. Dhingra, P. S. Soltis, R. Shaw, W. G. Farmerie, K. M. Folta, and D. E. Soltis. 2006. Rapid and accurate pyrosequencing of angiosperm plastid genomes. BMC Plant Biology 6: 17. [HTML] [PDF]
Moore, M. J., A. Tye, and R. K. Jansen. 2006. Patterns of long-distance dispersal in Tiquilia subg. Tiquilia (Boraginaceae): implications for the origins of amphitropical disjuncts and Galápagos Islands endemics. American Journal of Botany 93(8): 1163-1177. [HTML] [PDF]
Moore, M. J., and R. K. Jansen. 2006. Molecular evidence for the age, origin, and evolutionary history of the American desert plant genus Tiquilia (Boraginaceae). Molecular Phylogenetics and Evolution 39(3): 668-687. [HTML] [PDF]
Moore, M. J., J. Francisco-Ortega, A. Santos-Guerra, and R. K. Jansen. 2002. Chloroplast DNA evidence for the roles of island colonization and extinction in Tolpis (Asteraceae: Lactuceae). American Journal of Botany 89(3): 518-526. [HTML] [PDF]
Courses Taught
In the fall I teach the introductory Biology majors course, Organismal Biology (BIOL 100). This course introduces students to the basics of how plants and animals work, from the level of the genome all the way up to that of the whole organism. Along the way we compare and contrast how plants and animals deal with the basic needs of survival. As you might expect, I handle the plant parts of the course, whereas Ms. Cruz teaches the animal parts. Aside from the actual biology component of the course, BIOL 100 is designed to prepare students for the rigors of upper-level Biology and Neuroscience courses. If you have questions about the course, including whether or not you should take it if you have been exempted from the course due to AP credit, please do not hesitate to contact me.
If you are interested in learning more about plant diversity and/or about the systematic and phylogenetic methods we use to understand organismal diversity, you might consider taking my spring semester course, Plant Systematics (BIOL 323/324). Although the focus of the course is on flowering plants, the concepts and methods that students learn are applicable to most organisms. We cover a wide range of topics, including morphology, taxonomy, nomenclature, phylogenetic methods and theory, biogeography, speciation, hybridization, and character evolution. The lab portion of the course involves significant field and lab components. For example, students learn how to work with next-generation DNA sequencing data to reconstruct the evolution of plants, and spend some hands-on time learning about the phylogenetic diversity of plants. Once the long Oberlin winter breaks, we head outside and learn more about plant diversity, including important plants of the Ohio flora.
I also lead an upper-level Biology majors Seminar, Biogeography (BIOL 423), that is focused on exploring the causes and consequences of the geographic distributions of organisms through group discussions of recently published scientific journal articles. Students choose nearly all of the articles we read, and also pursue a writing-intensive course project. As a class, we also take an afternoon field trip to a local site of biogeographic interest; in 2014, we visited Castalia Prairie, which is the most intact tallgrass prairie left in Ohio.
I also teach a First-Year Seminar entitled The Sixth Extinction: Problems and Prospects in Biodiversity Conservation (FYSP 194). In this course we explore the challenges and opportunities surrounding the conservation of biodiversity using readings from a wide variety of sources. Among the topics we discuss are the value of biodiversity, the impacts of legislation and private property on biodiversity, the ramifications of ethical dilemmas in conservation management, the feasibility of saving species and restoring degraded habitat, and ways to encourage participation in biodiversity conservation. We also go on several weekend afternoon field trips to sites of local conservation interest. Some of the places we have visited in the past include Castalia Prairie (the best remaining example of tallgrass prairie in Ohio), Edison Woods (the largest remaining forest in northern Ohio), and NASA Plum Brook Station (which contains around 6000 acres of actively managed forest, prairie, and savanna).
Last updated on May 1, 2020
All images are the copyright of Michael J. Moore